Replication protein A (RPA) and HARP binding to DNA
Replication protein A (RPA) is the main eukaryotic single-stranded DNA binding protein, binding to ssDNA with high affinity and being abundant in the cell. RPA is involved in many aspects of cellular metabolism, including DNA replication, recombination and repair.
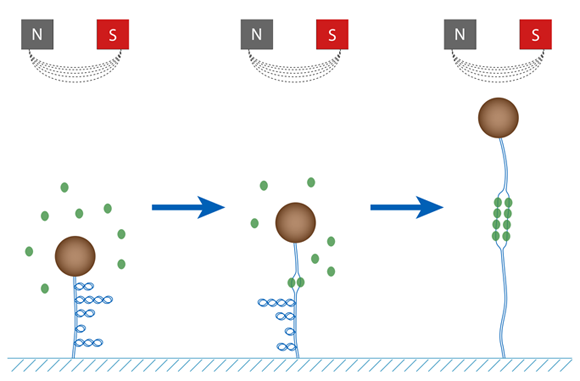
RPA is able to induce the unwinding of double-stranded DNA (dsDNA) by binding and stabilizing transiently formed bubbles of ssDNA (as seen in the figure below). Using magnetic tweezers we are able to study the dynamics of this reaction in real time and at the single molecule level.
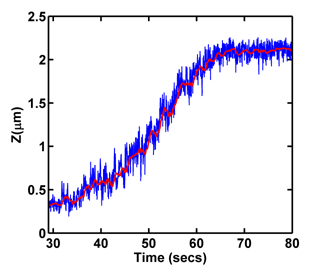
As seen in the figure monitoring magnetic sphere height, Z, we can infer details about the processes occurring on DNA. For example, we expect the sphere height to increase as RPA molecules bind to, and hence unwind, DNA. This is indeed observed experimentally as shown in the figure below.
Using this assay we find that unwinding rate is exponentially dependent on torsional stress present in the DNA. The unwinding reaction is self-limiting, ultimately removing the driving torsional stress. The RPA unwinding reaction serves to protect and stabilize the dsDNA when it is structurally destabilized by mechanical stress.
There are many dsDNA unwinding activities within cells and until recently no winding activity had been seen with which to balance it. Recently annealing helicases have been discovered with the unique ability to re-anneal un-wound dsDNA.
As dsDNA only transiently forms single stranded bubbles we create permanent stabilised bubbles which can then be acted upon by the annealing helicase. These experiments allows us to decipher the mechanism with which the proteins can re-anneal the stabilised ssDNA bubbles.
Reference:
Iwijn De Vlaminck, Iztok Vidic, Marijn T. J. van Loenhout, Roland Kanaar, Joyce H. G. Lebbink and, Cees Dekker, “Torsional regulation of hRPA-induced unwinding of double-stranded DNA”, Nucl. Acids Res. (2010) 38 (12): 4133-4142.