DNA supercoiling
In a cell, DNA is highly compacted to fit within its nucleus. If being stretched out, the DNA from all the cells in our body would reach Pluto. One way to compact DNA is by supercoiling: When rotations are applied to a DNA molecule, it can form higher-order structures, called DNA supercoils or plectonemes – see Figure 1.
DNA is constantly accessed and processed by enzymes such as RNA-, DNA-polymerases, topoisomerases, and helicases. Often, these enzymatic activities introduce torsional stress on the DNA, introducing or releasing plectonemes. These DNA supercoils, in turn, alter the activity of the DNA processing enzymes. Hence, understanding the structure and dynamics of DNA supercoil is prerequisite for comprehensive understanding of the enzymes and their biological functions.
Despite its fundamental importance, DNA supercoils and the associated biological processes are poorly understood, mainly due to the lack of proper experimental tools to examine them. Our laboratory has been pioneering in development of the techniques to visualize the supercoiled DNA at single-molecule level. We recently developed a novel technique to visualize the supercoiled DNA at single-molecule level (Fig.2, Ganji et al, Nano Lett, 2016). By using this technique, we studied how the formation of plectonemes is controlled by the DNA sequence and by local mechanical properties of the DNA (Kim et al, elife, 2018). Furthermore, we are currently studying the interactions of supercoiled DNA with proteins that generate, exploit, and remove DNA supercoils for their biological functions, such as RNA polymerase (Fig.3), nucleosome assembly proteins, and topoisomerase.
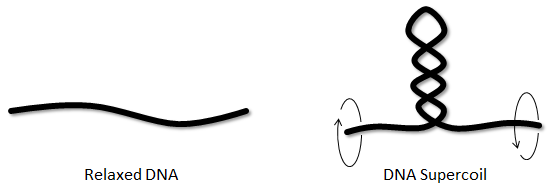
Figure 1. A coiled loop structure (plectoneme) can be formed when the DNA is twisted.
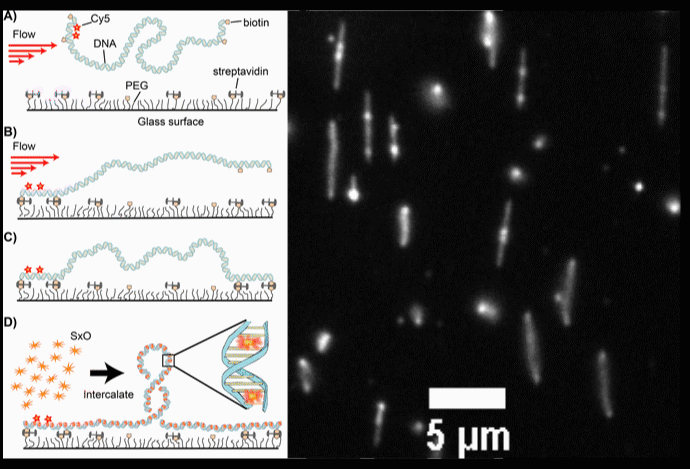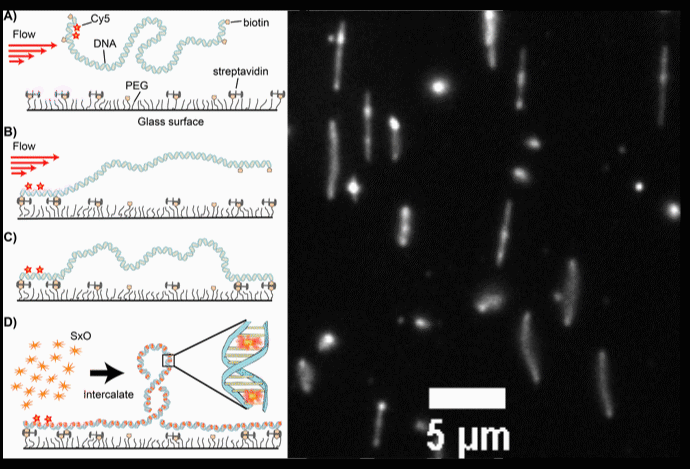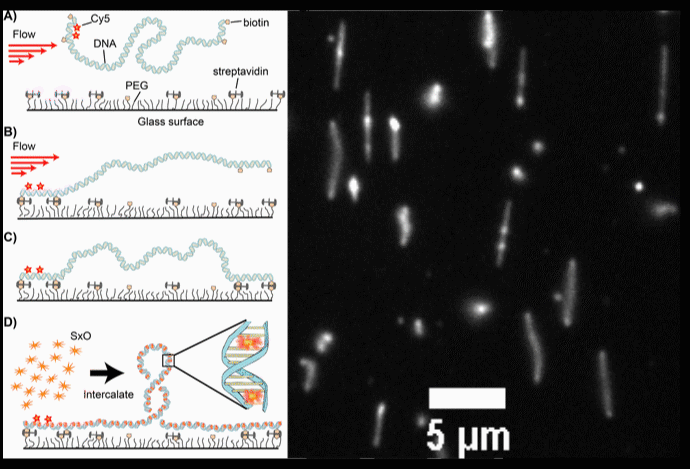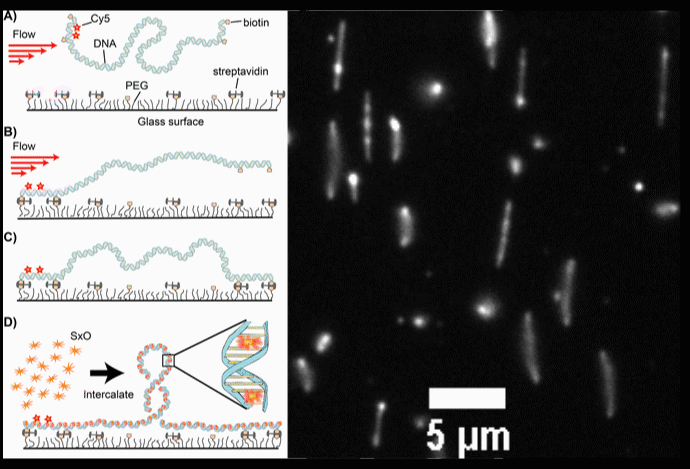
Fig. 2. (A-D) Schematic diagrams of the intercalation-induced Supercoiled DNA (ISD). (E) A fluorescence microscopy movie with multiple DNA molecules showing plectonemes as diffusing bright spots along DNA.
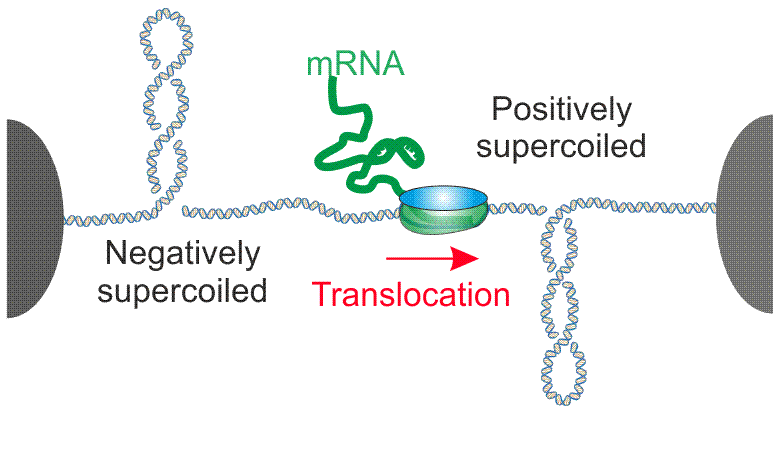
Fig. 3. Schematic of RNA polymerase –induced DNA supercoil formation.
People working on this project

Eugene Kim
Mahipal Ganji
Sung Hyun Kim

Jaco van der Torre
- Room F0.190
- +31-(0)15-27 83959
- J.vanderTorre@[REMOVE THIS]tudelft.nl
- Expertise: Molecular biology

Cees Dekker
- F0.210
- +31-(0)15-27 86094
- C.Dekker@[REMOVE THIS]tudelft.nl
- Principal Investigator
- View CV


